Why pre-analytical control matters for cardiovascular biomarker assays
Small handling differences can shift cytokine and soluble adhesion marker readouts more than biology does. That's why this serumplasma pre-analytical checklist focuses on time-to-spin, anticoagulant choice, platelet control, storage, and shipping discipline—so multi-site cohorts generate comparable data.
Three pillars keep variability in check:
- Standardize what you do and what you record. The SPREC framework encodes matrix, anticoagulant, pre-centrifugation time/temperature, processing type, and storage, enabling cross-site comparability (see the original standard by Betsou and colleagues on SPREC (2010)).
- Treat documentation like bioanalysis. Guidance such as FDA's Bioanalytical Method Validation (2018) and ICH M10 (2022) helps teams structure accuracy/precision/stability evidence under RUO framing.
- Lock matrix and processing windows before enrollment. Reviews on pre-analytics emphasize constrained timing and consistent centrifugation to reduce ex vivo drift across sites.
Project entry point for end-to-end execution: cardiovascular disease biomarker assay.
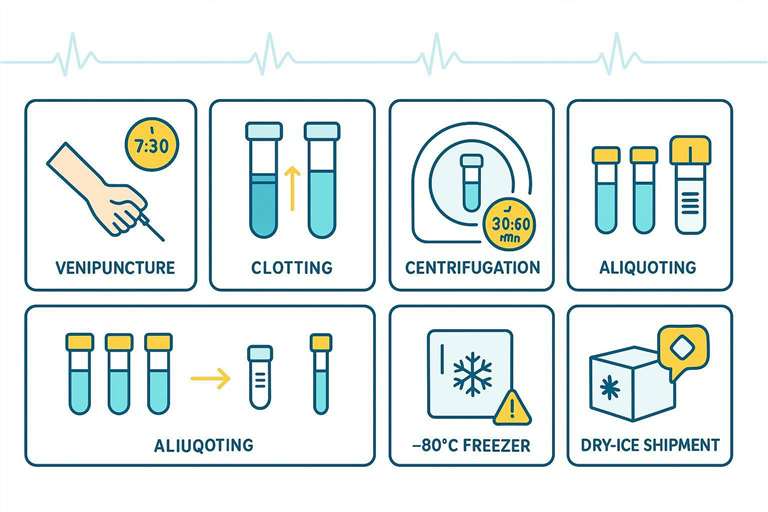 A standardized workflow to reduce site-driven artifacts in serum/plasma cardiovascular biomarker studies.
A standardized workflow to reduce site-driven artifacts in serum/plasma cardiovascular biomarker studies.
Serum vs plasma: choose the matrix that matches your study reality
Serum selection logic for cardiovascular biomarker assays
- The clotting step can affect certain analytes (e.g., platelet-derived factors); however, serum is often simpler for multi-site workflows because it avoids anticoagulant variability.
- For large cohorts, matrix consistency usually beats any small analytic advantage from a "perfect matrix."
- Literature comparing cytokines across anticoagulants shows differences that are modest yet meaningful longitudinally—for example, some cytokines trend lower in citrate plasma versus EDTA, underscoring the need to lock a single matrix/anticoagulant throughout the study, per findings summarized by Verberk et al. (2021).
Plasma selection logic for cardiovascular biomarker assays
- Anticoagulant matters: EDTA and citrate are the typical choices; avoid switching mid-study. Heparin can be acceptable for select targets but may not be preferred for some cytokine panels based on platform-specific behavior.
- When thrombo-inflammation markers (e.g., soluble CD40L, PF4, TGF-β1) are endpoints, control platelet activation: gentle inversion, prompt spin, and consider a validated second spin to generate platelet-poor plasma if required by your assay.
- For matrix planning and cross-matrix considerations, see serum & plasma cytokine assay.
Matrix decision table (fast pick)
| Study constraint | Prefer serum | Prefer plasma | Lock-in requirement |
|---|---|---|---|
| Many sites, simple SOP | ✓ | Single tube type across sites | |
| Platelet-sensitive markers included | ✓ | Tight processing timing + mixing rules | |
| Anticoagulant-specific protocols exist | ✓ | EDTA/heparin/citrate standardized | |
| Long storage with minimal handling | ✓ | ✓ | Aliquoting + freeze–thaw limits |
Collection checklist: tubes, timing, and site harmonization
Tube type and labeling controls
- Fix tube type per protocol and per site; if plasma is selected, specify EDTA or citrate and keep it constant.
- Apply barcoded IDs and capture timestamps for draw, first spin, and freeze; maintain a chain-of-custody across transfers.
- Map these fields to SPREC codes to aid cross-site reconciliation.
Time-to-spin and processing window
- Lock a processing window to prevent site drift. As starting windows (validate locally):
- Serum: allow 20–30 min clot at room temperature, then spin within 1–2 h of draw.
- Plasma: aim for first spin within 30–60 min; refrigerate if delays approach 2 h.
- Standardize centrifugation: 2,000–2,500 × g for 10–15 min (room temperature or 4°C if validated) with rotor radius, brake, and temperature documented. For platelet-poor plasma, add a second spin (e.g., 10,000–16,000 × g, ~10 min) per local validation, as operationalized in pre-analytical best-practice reports such as Gegner et al., 2022.
Site harmonization mini-audit
- Complete a training checklist at each site (phlebotomy, labeling, time-to-spin discipline, centrifuge use).
- Run a pilot batch to verify timing adherence and centrifuge calibration before full enrollment.
- Align deviation handling rules (out-of-window processing, temperature excursion, mislabeling) and record them consistently across sites.
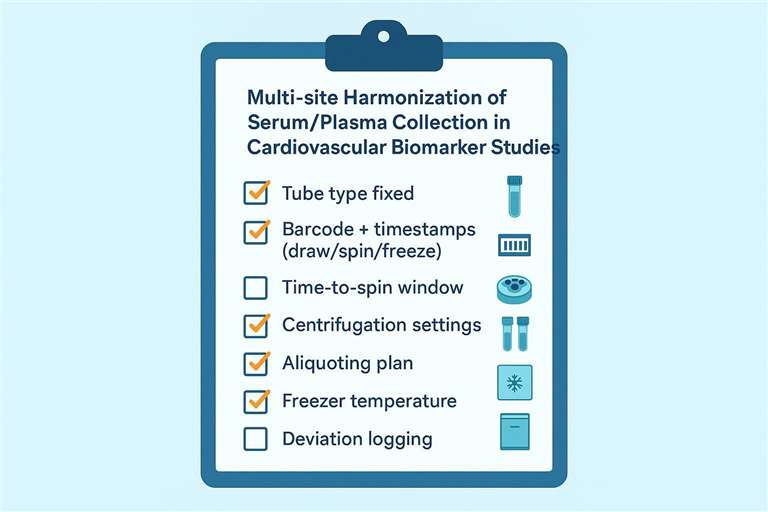 A site-level checklist to standardize collection and processing for cardiovascular biomarker assays.
A site-level checklist to standardize collection and processing for cardiovascular biomarker assays.
Aliquoting and storage checklist: protect stability and reduce freeze–thaw
Aliquot strategy for multiplex cardiovascular biomarker panels
- Use small, single-use aliquots to avoid repeated thawing; reserve replicate aliquots for QC and re-runs.
- Plan volumes against multiplex input needs and plate layouts to minimize bench time. For volume budgeting within xMAP multiplex workflows, see Luminex cytokine detection service.
Storage temperature and monitoring
- Lock long-term storage at −80°C with continuous monitoring and alarm policy.
- Document any temperature excursion (date/time, duration, min/max) and the disposition outcome; align sample maps to enable rapid retrieval without prolonged warming.
Freeze–thaw governance
- Track freeze–thaw count per aliquot and define rules by analyte class (accept/flag/exclude). A conservative RUO policy: design for 0 cycles; if reuse is unavoidable, cap at ≤2 (≤3 only with justification and flagging). Evidence reviews note heterogeneous and poorly documented effects across studies; transparency and local validation are essential, as summarized in Sjöbom et al. (2022).
Interference and rejection criteria: hemolysis, lipemia, icterus
Visual/metric screening for common interferents
- Record visual grading and, where available, instrument HIL indices to standardize decisions.
- Predefine per-assay sensitivity and thresholds via manufacturer guidance and local spike-in tests; methodology examples for interference assessment are reviewed by Ho et al., 2021.
Disposition rules (accept / flag / exclude / recollect)
- Set rules by analyte class (e.g., cytokines vs soluble adhesion markers) and platform.
- Distinguish "flag-only" from "exclude" thresholds and specify recollection triggers for critical timepoints.
Interference checklist table (field-ready)
| Interference type | Common cause | What to record | Disposition options |
|---|---|---|---|
| Hemolysis | Traumatic draw, delays | Visual grade / note | Accept with flag / exclude / recollect |
| Lipemia | Non-fasting, metabolic | Visual grade / note | Flag + dilution plan / exclude |
| Icterus | Hyperbilirubinemia | Visual grade / note | Flag + interpret cautiously / exclude |
Shipping checklist: cold chain, packaging, and documentation
Cold-chain requirements for serum/plasma shipments
- Plan dry ice mass for the route duration with safety margin; place dry ice between the secondary container and the insulated outer shipper.
- Use secondary containment with absorbent and ensure all primary receptacles are leakproof and sealed.
- Add a temperature logger for long or variable routes to provide objective evidence of cold-chain integrity—this supports consistent intake decisions for biospecimen shipments.
Packaging and compliance documentation
- Follow the triple-packaging system for Biological Substance, Category B (UN3373) as summarized by the CDC and carriers referencing IATA PI 650. Mark the outer package with the UN3373 mark and, when using dry ice (UN1845), add the Class 9 label and net dry ice weight.
- Include a sample manifest that reconciles aliquot counts with barcodes. No Shipper's Declaration for Dangerous Goods is required for UN3373; use an Air Waybill or equivalent that notes UN3373.
- On receipt, check for damage, thaw evidence, and review any logger data against acceptance criteria. Practical job aids and examples are shown in CDC's Category B labeling and documentation job aid.
Shipping handoff to assay execution
- At intake, align sample identifiers, matrix, and freeze–thaw metadata with the assay worklist for cardiovascular disease biomarker assay.
- Ensure matrix notes and anticoagulant details match expectations for the planned panel; cross-reference serum & plasma cytokine assay.
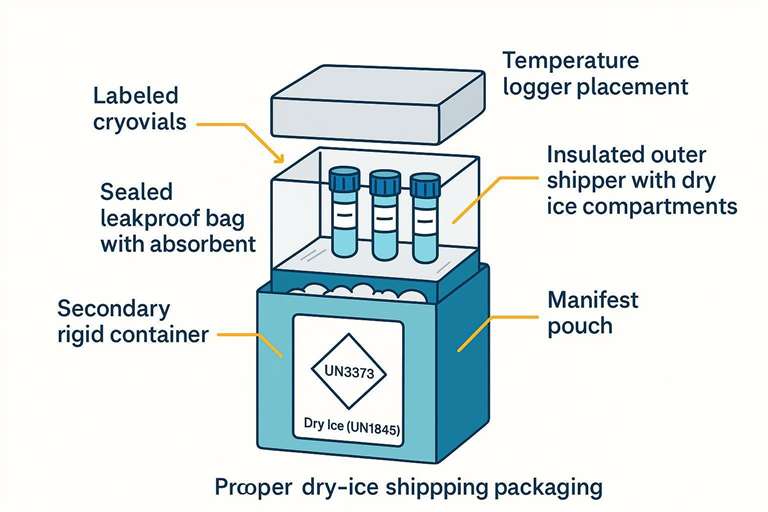 Packaging layers and documentation to preserve cold chain during shipment.
Packaging layers and documentation to preserve cold chain during shipment.
Reporting checklist: what to include with every sample batch
Minimum metadata bundle
- Sample matrix and tube type (serum/plasma; anticoagulant).
- Draw/spin/freeze timestamps and adherence to the processing window; centrifugation settings (g, time, temperature).
- Freeze–thaw count per aliquot and storage temperature.
Batch documentation bundle
- Site ID and collection operator ID (where applicable).
- Deviation log with corrective actions and disposition decisions.
- Shipping details: courier, dry ice weight, logger result summary (if used).
- Align report fields with LLD/LLOQ/ULOQ definitions to streamline analysis-stage QA; for background, see Understanding LLD, LLOQ, and ULOQ in Luminex Cytokine Assays.
Related cluster articles
- Platform and cohort scaling context: How to Choose a Multiplex Cardiovascular Biomarker Assay for Cohort Studies
- QC acceptance rules (analysis-stage): Multiplex Biomarker Assay QA/QC Acceptance Criteria
- Panel planning context: How to Design a Cardiovascular Biomarker Panel
FAQ
Serum vs plasma: which is better for cardiovascular biomarker assays?
Either can work under RUO conditions. The better choice is the one your sites can standardize with tight timing and a fixed anticoagulant, especially if platelet-sensitive markers are included.
How many freeze–thaw cycles are acceptable for cytokines and soluble biomarkers?
Design for single-use aliquots. If reuse is unavoidable, cap at ≤2 cycles (≤3 only with justification and flags) and document counts per aliquot.
What should I do if a sample is hemolyzed or lipemic?
Record the interference (visual grade/index), apply pre-set rules (accept with flag, dilute/clarify if validated, exclude, or recollect), and keep decisions consistent across sites.
How do I standardize sample collection across multiple sites?
Lock the SOP, train and verify with a pilot batch, barcode and timestamp every handoff, and use a clear deviation taxonomy with consistent disposition rules.
Do I need a temperature logger for dry-ice shipments?
Use a logger when routes are long or variable; it provides objective evidence of cold-chain integrity to support consistent intake decisions.
References:
- U.S. Food and Drug Administration (FDA). (2018). Bioanalytical Method Validation: Guidance for Industry [PDF].
- European Medicines Agency (EMA) / ICH. (2022). ICH Guideline M10 on Bioanalytical Method Validation and Study Sample Analysis (Step 5) [PDF].
- Betsou, F., Lehmann, S., Ashton, G., et al. (2010). Standard preanalytical coding for biospecimens: Defining the Sample PREanalytical Code (SPREC). Cancer Epidemiology, Biomarkers & Prevention, 19(4), 1004–1011. https://doi.org/10.1158/1055-9965.EPI-09-1268
- Verberk, I. M. W., et al. (2021). Pre-analytical sample handling effects on blood cytokine levels: Quality control of a COVID-19 biobank. Bioanalysis. https://doi.org/10.2217/bmm-2020-0770
- Gegner, H. M., et al. (2022). Pre-analytical processing of plasma and serum samples for combined proteome and metabolome analysis. Frontiers in Molecular Biosciences. https://doi.org/10.3389/fmolb.2022.961448
- Sjöbom, U., Nilsson, A. K., Gyllensten, H., Hellström, A., & Löfqvist, C. (2022). A systematic review and meta-analysis of preanalytical factors and methodological differences influencing the measurement of circulating vascular endothelial growth factor. PLOS ONE, 17(7), e0270232. https://doi.org/10.1371/journal.pone.0270232
- Centers for Disease Control and Prevention (CDC). (2022). Step 4: Labeling, Marking, and Documenting Requirements Job Aid (Biological Substance, Category B [UN3373]) [PDF].

